看了null的用户又看了
日本RIBM HS-AFM
超高速视频原子力显微镜
| 产品简介:
超高速视频原子力显微镜(Sample-Scanning High-Speed Atomic Force Microscope ,HS-AFM SS-NEX)是由日本 Kanazawa 大学 Prof. Ando 教授团队研发的,也是上台可以达到视频成像的商业化原子力显微镜。HS-AFM突破了传统原子力显微镜“扫描成像速慢”的限制,能够实现在液体环境下超快速动态成像,分辨率为纳米水平。样品无需特殊固定,不影响生物分子的活性,尤其适用于生物大分子互作动态观测。推出至今,全球已有80多位用户,发表 SCI 文章 200 余篇,包括Science, Nature, Cell 等杂志。 |
技术特征:
| 扫描速度快 | 探针小,适用于生物样品 | 自动校正,适合长时间测样 |
| ◆ 扫描速度高可达 20 frame/s ◆ 有 4 种扫描台可供选择 | ◆ 悬臂探针共振频率高,弹簧系 数小。避免了对生物样品等的损伤 | ◆ 悬臂探针可自动漂移校准, 适用于长时间观测。 |
技术原理:
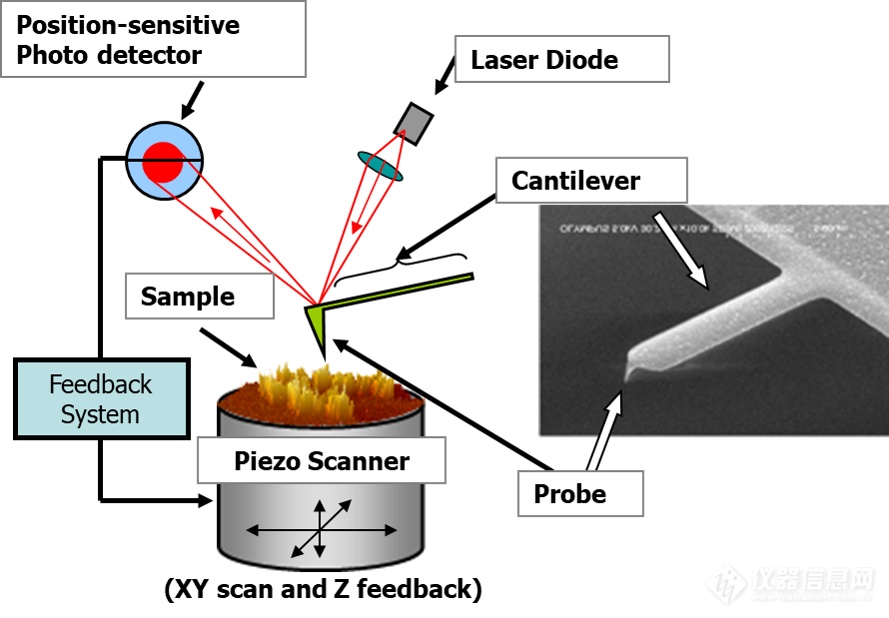
超高速视频原子力显微镜应用领域:
应用包括:利用
HS-AFM可在纳米尺度动态实时记录生物大分子的运动以及分子间相互作用,包括:
|
|
|
|
walking myosin V 实时观察 | dendrite growth in neuron 实时观察 | rotorless F1-ATPase 实时观察 | light response for D69N 实时观察 |
 |  |  |  |
| IgG antibody 150nm * 150nm | plasmid DNA 250nm * 250nm | myosinⅡ 500nm * 500nm | bacteriorhodopsin 40nm * 40nm |
 |  |  |  |
lipid membrane 3500nm * 3500nm | 350nm poly beads 900nm * 900nm | E.coli 3000nm * 3000nm | 350nm poly beads 3000nm * 3000nm |
超高速视频原子力显微镜相关应用案例:
1.Video imaging of walking myosin V 实时观察myosin V蛋白的运动
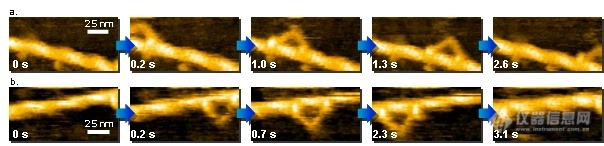

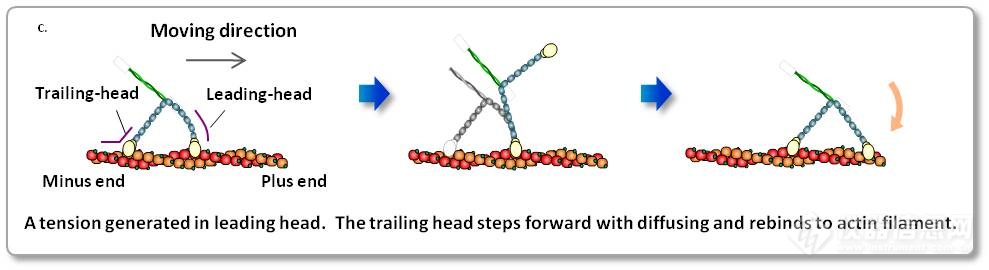
N. Kodera et al. Nature 468, 72 (2010). Kanazawa University
2.Real-space and real-time dynamics of CRISPR-Cas9 实时显示CRISPR基因编辑
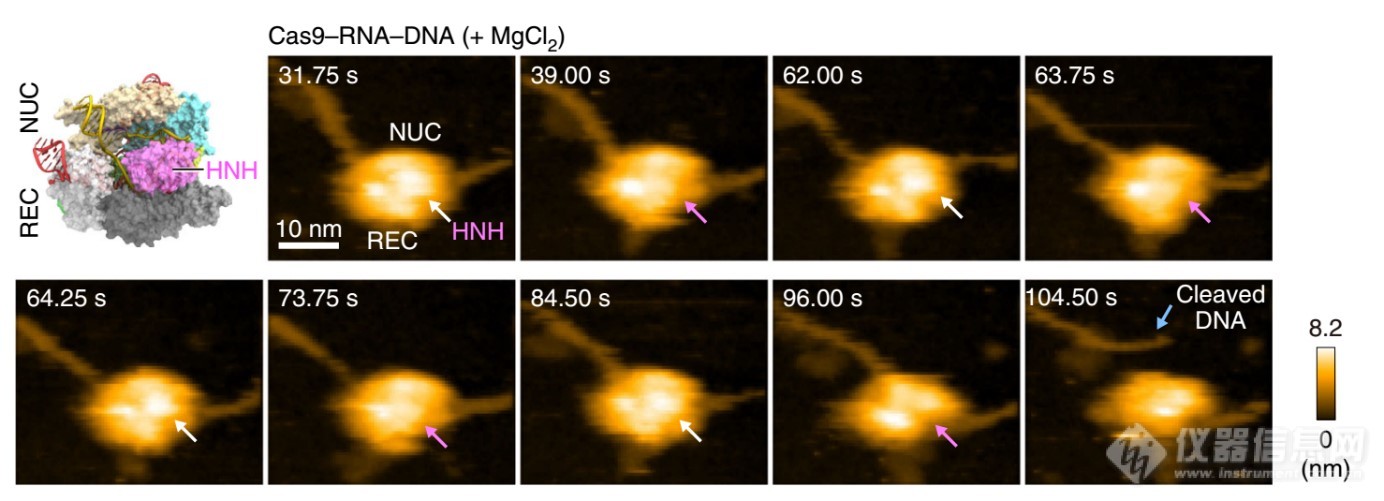
Mikihiro et al. Nature Communications, (2017). Kanazawa University
设备规格及参数:
| 标准配置 | |
| 扫描速度 scan speed | 50 ms/frame (20 frames/sec) |
| 压电扫描器 piezo range | X: 0.7μm, Y:0.7μm, Z: 0.4μm |
| 样品大小 sample size | 1.5mm in diameter |
| 扫描环境 environment | In liquid/In air |
| 控制系统 control system | PID control, Dynamic PID control |
| significant Function | Scanner active dumping,Drift correction for cantilever excitation |
| 可选配置 | |
光照装置 Light irradiation Unit | Light irradiation unit for the experiments with caged compounds. Variable wavelength: 350-560nm
|
宽扫描台 wide scanner | 1frames/s;XY:4μm×4μm, Z:0.7μm |
超宽扫描台Amplified ultra wider scanner | 0.1frames/s;XY:30μm×30μm, Z:1.2μm |
微流控系统 Circulation unit | The observation solutions can be exchanged while continuing AFM observation. |
已发表文献(2017年):
1. Ando T.; "Directly watching biomolecules in action by
high-speed atomic force microscopy"; Biophys. Rev.
(2017)
2. Ando T.; "High-speed Atomic Force Microscopy for
Observing Protein Molecules in Dynamic Action", Proceedings of SPIE
10328, Selected Papers from the 31st International Congress on
High-Speed Imaging and Photonics (2017)
3. Aybeke E., Belliot G., Lemaire‐Ewing S., Estienney M.,
Lacroute Y., Pothier P., Bourillot E., Lesniewska, E.; "HS‐AFM and SERS Analysis
of Murine Norovirus Infection: Involvement of the Lipid Rafts"; Small 13 1 (2017)
4. Cai W, Liu Z., Chen Y., Shang G.; "A Mini Review of the
Key Components used for the Development of High-Speed Atomic Force Microscopy";
Science of Advanced Materials Vol. 9 Numb. 1 (2017) p.77-88
5. Colom A., Redondo-Morata L., Chiaruttini N., Roux A., Scheuring S.; "Dynamic remodeling of the dynamin helix during membrane constriction"; Proceedings of the National Academy of Sciences 114 21 (2017)
6. Dufrêne Y., Ando T., Garcia R., Alsteens D.,
Martinez-Martin D., Engel A., Gerber Ch., Müller D.; "Imaging modes of atomic
force microscopy for application of molecular and cell biology"; Nat.
Nanotechnol. 12 (2017) p.295-307
7. Harada H., Onoda A., Uchihashi T., Watanabe H.,
Sunagawa N., Samejima M., Igarashi K., Hayashi T.; "Interdomain flip-flop motion
visualized in flavocytochrome cellobiose dehydrogenase using high-speed atomic
force microscopy during catalysis"; Chemical Science (2017)
8. Karner A., Nimmervoll B., Plochberger B., Klotzsch E.,
Horner A., Knyazev D., Kuttner R., Winkler K., Winter L., Siligan Ch., Ollinger
N., Pohl P., Preiner J.; "Tuning membrane protein mobility by confinement into
nanodomains"; Nature Nanotechnology 12 3 (2017)
p.260-266
9. Keya J., Inoue D., Suzuki Y., Kozai T., Ishikuro D.,
Kodera N., Uchihashi T., Kabir A., Endo M., Sada K., Kakugo A.; "High-Resolution
Imaging of a Single Gliding Protofilament of Tubulins by HS-AFM" ;
Scientific Reports 7 1 (2017)
10. Kim Y.; "An Advanced Characterization Method for the
Elastic Modulus of Nanoscale Thin-Films Using a High-Frequency Micromechanical
Resonator"; Materials 10 7 (2017)
11. Kim Y.; "An evaluation technique for high-frequency
dynamic behavior of a sandwich microcantilever beam"; Journal of
Sandwich Structures & Materials (2017)
12. Korolkov V., Baldoni M., Watanabe K., Taniguchi T., Besley E., Beton P.; "Supramolecular heterostructures formed by sequential epitaxial deposition of two-dimensional hydrogen-bonded arrays"; Nature Chemistry (2017)
13. Legrand B., Salvetat J.-P., Walter B., Faucher M.,
Théron D., Aimé J.-P.; "Multi-MHz micro-electro-mechanical sensors for atomic
force microscopy"; Ultramicroscopy 175 (2017) p.46-57
14. Liao H.-S., Chih-Wen Yang, Hsien-Chen Ko, En-Te Hwu, Ing-Shouh Hwang; "Imaging initial formation processes of nanobubbles at the graphite–water interface through high-speed atomic force microscopy"; Applied Surface Science (2017)
15. Matsui S., Kureha T., Hiroshige S., Shibata M.,
Uchihashi T., Suzuki D.; "Fast Adsorption of Soft Hydrogel Microspheres on Solid
Surfaces in Aqueous Solution"; Angewandte Chemie (2017)
16. Mierzwa B., Chiaruttini N., Redondo-Morata L., Moser von Filseck J., K?nig J., Larios J., Poser I., Müller-Reichert T., Scheuring S., Roux A., Gerlich D.; "Dynamic subunit turnover in ESCRT-III assemblies is regulated by Vps4 to mediate membrane remodeling during cytokinesis"; Nature Cell Biology (2017)
17. Miyata K., Tracey J., Miyazawa K., Haapasilta V., Spijker P., Kawagoe Y., Foster A., Tsukamoto K., Fukuma T.; "Dissolution Processes at Step Edges of Calcite in Water Investigated by High-Speed Frequency Modulation Atomic Force Microscopy and Simulation"; Nano Lett. 17 7 (2017) p.4083-4089
18. Miyazawa K., Watkins M., Shluger A., Fukuma T.; "Influence of ions on two-dimensional and three-dimensional atomic force microscopy at fluorite–water interfaces"; Nanotechnology Vol. 28 Numb. 24 (2017)
19. Mohamed M., Kobayashi A., Taoka A., Watanabe-Nakayama T., Kikuchi Y., Hazawa M., Minamoto T., Fukumori Y., Kodera N., Uchihashi T., Ando T., Wong R.; "High-Speed Atomic Force Microscopy Reveals Loss of Nuclear Pore Resilience as a Dying Code in Colorectal Cancer Cells"; ACS Nano 11 6 (2017) p.5567-5578
20. Nievergelt A., Andany S., Adams J., Hannebelle M., Fantner G.; "Components for high-speed atomic force microscopy optimized for low phase-lag"; Proceedings of 2017 IEEE International Conference on Advanced Intelligent Mechatronics (AIM) (2017)
21. Rangl M., Rima L., Klement J., Miyagi A., Keller S., Scheuring S.; "Real-time Visualization of Phospholipid Degradation by Outer Membrane Phospholipase A using High-Speed Atomic Force Microscopy"; Journal of Molecular Biology 429 7 (2017) p.977-986
22. Ren J., Zou Q.; "High-speed dynamic-mode atomic force microscopy imaging of polymers: an adaptive multiloop-mode approach"; Beilstein J. Nanotechnol. 8 (2017) p.1563-1570
23. Ricci M., Trewby W., Cafolla C., Vo?tchovsky K.; "Direct observation of the dynamics of single metal ions at the interface with solids in aqueous solutions"; Scientific Reports 7 43234 (2017)
24. Rigato A., Miyagi A., Scheuring S., Rico F.; "High-frequency microrheology reveals cytoskeleton dynamics in living cells"; Nature Physics (2017) DOI: 10.1038/NPHYS4104
25. Ruan Y., Miyagi A., Wang X., Chami M., Boudker O.,
Scheuring S.; "Direct visualization of glutamate transporter elevator mechanism
by high-speed AFM"; PNAS 114 7 (2017) p.1584-1588
26. Sadeghian H., Herfst R., Dekker B., Winters J., Bijnagte T., Rijnbeek R.; "High-throughput atomic force microscopes operating in parallel"; Review of Scientific Instruments 88 033703 (2017)
27. Sakiyama Y., Panatala R., Lim R.; "Structural Dynamics of the Nuclear Pore Complex"; Seminars in Cell and Developmental Biology (2017)
28. Shibata M., Watanabe H., Uchihashi T., Ando T., Yasuda R.; "High-speed atomic force microscopy imaging of live mammalian cells"; Biophysics and Physicobiology Vol. 14 (2017) p.127-135
29. Terahara N., Kodera N., Uchihashi T., Ando T., Namba K., Minamino T.; "Na+-induced structural transition of MotPS for stator assembly of the Bacillus flagellar motor"; Science Advances 3 11 eaao4119 (2017)
30. Uchihashi T., Scheuring S.; "Applications of high-speed atomic force microscopy to real-time visualization of dynamic biomolecular processes"; Biochim Biophys Acta. (2017)
31. Usukura E., Narita A., Yagi A., Sakai N., Uekusa Y.,
Imaoka Y., Ito S., Usukura J.; "A Cryosectioning Technique for the Observation
of Intracellular Structures and Immunocytochemistry of Tissues in Atomic Force
Microscopy (AFM)"; Scientific Reports 7 (2017)
32. Watanabe S., Ando T.; "High-speed XYZ-nanopositioner for scanning ion conductance microscopy"; Applied Physics Letters 111 11 (2017)
33. Watanabe-Nakayama T., Kodera N., Konno H., Ono K., Teplow D., Yamada M., Ando T.; "Nano-Space Video Imaging Reveals Structural Dynamics of Fibrous Protein Assembly and Relevant Enzymes"; Biophysical Journal 112 3 (2017)
34. Zhang Y., Tunuguntla R., Choi P., Noy A.; "Real-time dynamics of carbon nanotube porins in supported lipid membranes visualized by high-speed atomic force microscopy"; Philosophical Transactions of The Royal Society B Biological Sciences 372 (2017)
35. Zhang Y., Yoshida A., Sakai N., Uekusa Y., Kumeta M., Yoshimura S.; "In vivo dynamics of the cortical actin network revealed by fast-scanning atomic force microscopy" Microscopy 20 (2017) p.272-282
部分用户列表:

最多添加5台
